By: Jim Brewster
From the discovery of cells to the mapping of the human genome, let’s celebrate DNA Day by tracing the remarkable scientific progress that unlocked the secrets of genetics.
Ah, DNA Day. The day when we celebrate, well, DNA. At first glance, you may be rather underwhelmed by such a commonplace substance.
Why have a whole day devoted to spirals? You do that every time you eat rotini, right?
April 25 is the day in 1953 when a paper published by James Watson and Francis Crick definitively described the structure of DNA as a double helix chain. DNA Day celebrates this achievement, but to me, what it really celebrates is the monumental advances in science that brought us the understanding we have today of the fundamental building blocks of life.
The iconic double helix is ubiquitous in our society, but when you step back and think about how we as a species were able to discover the intricate beauty of something so tiny, it really boggles the mind.
The average human is made of 36 trillion cells. In each cell is a nucleus, and in each nucleus is DNA. The DNA weighs less than six picograms. How much is a picogram? I have no idea, but after a bit of Googling, I discovered that the mathematical formula for it is one divided by a lot. Not only is it tiny, but it is composed of even smaller bits of information that are vital to the proper function of life.
Determining the information inside a chromosome, inside a nucleus, inside a cell, and inside your body is known by many as genetics and by literally no one as in-cell-ption. I am hoping incellption catches on, though, since it is a way punnier name.
As a species, how did we dig out this information and determine the elegant double helix structure we all now recognize?
How did we go from realizing a cell exists to recognizing the entire human genome?
Come along, friends and neighbors, as we start at the top layer of the incellption and work our way down. This journey has it all: words, punctuation, and even the occasional picture!
Unveiling the microscopic world and the discovery of the cell
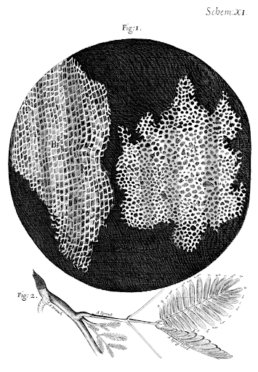
Today, this term is best known as the sole contents of an orange cat’s head, but the term comes from a very different source. In 1665, Robert Hooke refined the microscope with multiple lenses to produce a magnification of 200x. This allowed the study of “microscopic” organisms for the first time. After observing cork under his newfangled contraption, he saw structures that reminded him of the living quarters of monks (the original meaning of the word cell). Thus the name was coined in his succinctly titled, Micrographia: or Some Physiological Descriptions of Minute Bodies Made by Magnifying Glasses. With Observations and Inquiries Thereupon.
Shortly after the publication of Hooke’s, I Couldn’t Think of a Shorter Title, the Dutch scientist Antonie van Leeuwenhoek picked up a microscope and started examining bacteria and microbes for the first time. He saw these tiny “animals” wiggling around, living their best microscopic lives, and named them “animalcules.” It became clear that these animalcules were present in people, too, but what exactly were they? How did they reproduce? How did they feel about pineapple on pizza? These questions and more would remain unanswered for quite some time.
Unraveling the enigma of the nucleus
Since cells are teeny tiny animals, they have teeny tiny organs called organelles. The first of these to be discovered was the nucleus. Our buddy van Leeuwenhoek first described them in the red blood cells of salmon and named them “lumen.” It wouldn’t get the name we all know today until 1831 when a Scottish botanist, Robert Brown, called this opaque area (which was all the resolution of the day would show) the “nucleus.” The function of the nucleus, however, remained unclear for quite some time. A series of experiments in the mid-1800s demonstrated it played some role in heredity and that it was essential to cell formation, but the mystery of what it actually did would have to wait until the next layer of incellption that was uncovered in the late 1800s and the first years of the 20th century.
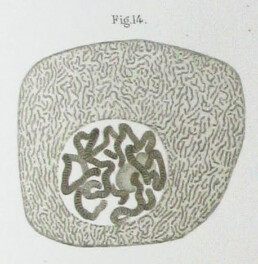
Painting a picture of inheritance through chromosomes
Advances in magnification and cell staining led to the discovery of structures within the nucleus. Cell staining is adding a dye to a culture to better identify the structures within. This lends the chromosome its name, as it combines the Greek words for color (chroma) and body (soma). This ability for chromosomes to absorb dye was later observed by Paul Simon, who described it in his 1973 classic hit “Chromosome.”
“Chromosome
Give us those nice bright colors
Give us the greens of summer
Make you think all the world is a celly day oh yeah”
At least I think that’s how the song went. The same method is widely used today and even used to add clarity to CT scan images. The ability to clearly discern the shape of chromosomes was important for recognizing them as the key factor in inheritance. By identifying the shapes and number of chromosomes, it definitively proved that chromosomes were inherited in equal numbers by both parents and, therefore, a key factor in heredity. This led to the final level of incellption, and the actual focus of this article: DNA.
DNA’s structural revelation
The seminal paper published by Watson and Crick in April 1953 that described the structure of DNA was actually a culmination of work done by several other scientists on the chemical properties and helix shape of DNA. This can be summarized in three sections: Nucleic acid, nucleotides, and helix shape.
Nucleic acid
In the late 1800s, a Swiss physician, Johannes Miesher, tested the contents of the newly discovered nucleus and found it to be an acidic mixture that was high in phosphorus and nitrogen. He called this mixture “nuclein,” which was later renamed “nucleic acid” due to its high acidity. Personally, I want to live in the alternate reality where animalcules are powered by nuclein, because that just sounds cooler, but oh well.
Later studies showed the nucleic acid to include polymer chains held together with sugar called deoxyribose. Thus the mixture was called deoxyribose nucleic acid, or, as we more commonly know it, DNA.
DNA was also found to contain four specific chemicals: adenine, guanine, thymine, and cytosine. These are rather big words (unlike all the other jargon in this article, right?), so we will just refer to them by their first letters: A, G, T, and C. Today, we know these as “nucleotides” and understand their significance, but for a long time their pattern wasn’t recognized. Nucleotides were thought to be just another part of the homogenous nucleic acid and thus incapable of containing the information needed for hereditary traits. It was written off as a curiosity and attention was turned to proteins as the building blocks for quite some time.
Nucleotides
In the 1940s, a rather clever biochemist named Erwin Chargaff began studying nucleic acid using a technique called chromatography. This is a way to separate the components of a mixture to measure their amounts.
Chargaff began to see that the amount of each of the four nucleotides were not random, or homogenous, but tended to be consistent within a species. It might vary from one species to another, but within a species it was always the same, and not only that, but the amount of T was always roughly equivalent to the amount of A, and the same was true for G and C.
For example, if a nucleus contained 20% T, It also had 20% A. The remaining 60% was split evenly between G and C. The significance of this was that the mixture was not homogenous as previously thought, and this brought attention back to DNA as the candidate for the information passed along in heredity.
Helix shape
At the same time Watson and Crick were investigating the structure of DNA, Rosiland Franklin and Maurice Wilkins were exploring the shape of chromosomes using X-rays. X-rays shoot through an object, bounce around, and reflect back. Things with different densities will bounce around at different rates, and with this we can tell the internal structure of an object. This is how the dentist is able to discover all the cavities in your mouth, because teeth have a different density than the lack of teeth.
Franklin and Wilkins used this method to discover the structure of the chromosome known to reside in the nucleus thanks to earlier chromatographic pictures and stains. In the same 1953 edition of Nature, where Watson and Crick published their study on the structure of DNA, Franklin and Wilkins each published papers on the internal structure of chromosomes using X-ray imaging. Together, these papers demonstrated that DNA was made of a helical (spiraling) polymer chain, with evenly spaced, two chain nucleotides attached to it horizontally. In other words, it was a spiral chain of phosphate “backbone” with nucleotide pairs running all along it.
However, the number of nucleotides on each “row” was surmised based on the width of the spiral and the number of molecules that could fit side by side inside such a spiral. They guessed that there must be more than one intertwined polymer chain, but the number of chains and the way they connected to the nucleotides was anyone’s guess.
Double helix structure
Enter James Watson and Francis Crick. Working at Cambridge University, these two scientists sat down with the different lines of evidence we had so far about DNA:
- Deoxyribose nucleic acid is the main component.
- The ribose is made up of spiraling chains of phosphate and nitrogen.
- Attached to the chain are pairs of the nucleotides A, C, T and G.
- The amounts of A and T are equal, and the amounts of G and C are equal.
So, what can we do with this information? The two made ball-and-stick models of the molecules involved and found that A and T just fit together, and G and C just fit together. Fitting means they don’t get in each other’s way, but nothing is holding them together, either. However, A doesn’t fit with G or C, and G and C don’t fit with T. If each row has two nucleotides, and A and T are in equal amounts, then it follows that each row is a fitted AT pair. The same logic holds true for GC pairs. Now we have the pairs identified on each row. So far, so good.
Next, how do they fit to the backbone? Earlier models suggested a central core of phosphate with the nucleotides sticking off the sides. All four can bind to the backbone well. Binding means they share a hydrogen atom and make the atoms inseparable. That meant that if A was attached to the backbone, it wasn’t going anywhere, but nothing was stopping T from falling right off A on the other side. Not a great setup if you want something that will survive generations without parts falling off all over the place.
Instead, they tried making a model with two phosphate chains (because, remember, Franklin and Wilkins said there were probably more than one) that had the nucleotide pairs sandwiched in between them. Voila! The space was just enough to hold the nucleotides in place, but they were not bound. If you unwound the two strands, each nucleotide would stay exactly where it was supposed to be, and when you wound it back together, it would be right next to its partner. In their paper, they even noted, “It has not escaped our notice that the specific pairing we have postulated immediately suggests a possible copying mechanism for the genetic material.”
Think about it: DNA needs to make copies of itself to pass along to other cells and the next generation. The nucleotides, fitting but not binding, allow the DNA to open up to be copied, like opening a book to the page you want and pressing it against a photocopier. This is basically what RNA does, by the way. If A and T always pair together, then you could take just one of the two strands and know exactly what is on the other strand. Everywhere you see an A on one, you know there is a T on the other one, even without looking at it. This would be a perfect way to synthesize new DNA.
DNA Day
As mentioned earlier, April 25, 1953 was the day the article explaining this structure was published. This one-page, elegant solution to the problem of heredity was the culmination of literally centuries of work. It opened the door to the information storehouse of how the body functions and what is needed to create life.
Why did it take me approximately one eternity to get to this point in the article? Why didn’t I just say that up front? Well I did say that this article had words and sentences and whatnot in it. You were warned. No backsies.
So many intuitive leaps went into each stage of the journey. The helix shape, with its intricate mosaic of nucleotides, didn’t just jump out at the first person to look under a microscope. It took multiple lines of evidence gleaned from a variety of techniques and sources over lifetimes.
Watson and Crick are credited with “discovering” the structure, but they stood on the shoulders of so many giants to accomplish this. I think there is so much to celebrate in this 1953 achievement. To me, the paper might as well be sung to the tune of the Irish folk song Rattlin’ Bog:
And in this pair there was a nucleotide
A rare nucleotide, a rattlin’ nucleotide
And the nucleotide in the pair
And the pair in the helix
And the helix in the chromosome
And the chromosome in the nucleus
And the nucleus in the cell
And the Cell in the scientist
And the scientist in the lab
And the lab in the building
And the building down in the valley-o
The following section, mapping the genome, takes this human pyramid of genius one step further as we make more intuitive leaps full of more big words.
Mapping the genome
At this point in our story, we as a species knew that the code for life of every species is held in their DNA. Scientists quickly began peeling apart the helix to see what nucleotides lie within, and found an amazingly long, complex chain of nucleotides. Applications for inherited diseases and cancer research were immediately apparent. Lots of information was clearly there, but it was not simply just a long stretch of ACTG. It was an alternating sequence of all four in a seemingly random pattern. Many portions of the pattern were the same in everyone, but some sections of the genome seemed to vary between individuals. It was so large and varied that it was difficult to get a systematic list of differences without a point of reference. If there was a deviation from the standard, what was the standard? Having a standard map of the genome would help to identify differences in populations (e.g., breast cancer patients vs. prostate cancer patients) and what they meant.
In 1985 a workshop to start work on just such a reference sequence was held. This was projected to take about 15 years and in the end took just under 18. The Human Genome Project, as it became called, was an international effort from the governments of several countries, and included dozens of labs and scientists. To understand what a monumental achievement this is, we need to take a step back to appreciate the extreme size of the genome.
If you unwound DNA and laid it out in a straight line, it would measure approximately two meters. Two meters of microscopic nucleotides that number just over 3 billion base pairs. Three. Billion. Even using fingers and toes, I could do about 20, max. These base pairs are distributed along 13 chromosome pairs, so even if there was a way to unravel each one all at once, it is not feasible to measure all three billion bases at once. They decided to look at a little piece at a time instead of using an approach called “Hierarchical shotgun sequencing.”
Piecing together the genetic puzzle
All types of sequencing work by chopping DNA into smaller fragments that can be read. Hierarchical sequencing works by using “restriction enzymes” that cut the DNA at specific points, with fragments being at most 800 base pairs. When you cut 3 billion base pairs into 800 chunk fragments, you wind up with a lot. I’m not sure how many; I lost count at 20, but it’s a lot. It’s hard to tell how these pieces fit together, though.
It’s like dumping out a puzzle on a table. You know you have all the pieces there, and a general idea of what your end result will look like, but it will take some time to fit all the pieces together. Not only that, but each fragment isn’t cut all at once in even cuts. There is overlap with all of them. So it’s like a puzzle where the pieces more or less fit, but some overlap. You have to set them on top of each other so they line up. Add to this the fact that a ton of the pieces all look the same because DNA repeats itself a lot. It’s like the puzzle is like a picture of a sky that is mostly blue with some clouds here and there.
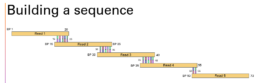
The first 10 years of the project were spent identifying landmark regions. These distinct areas can act as anchor points to attach the sea of repeating fragments. Continuing our puzzle analogy, it’s like identifying the clouds so you can piece together the overlapping blue sky pieces connected to the clouds and work your way out from there. To illustrate the overlap, take a look at my diagram below.
Let’s say Read 1 is our landmark region. We know this unique section ends in a sequence of ACTG. Next, we find a fragment that matches that ATCG section so we can fit it in place. When we continue this out, we can build a continuous read made of smaller, overlapping fragments. This painstaking process was done largely by hand. Shortly after the landmark regions were identified and the process of picking up the pieces began, another privately owned company called Celera stepped in with a different approach. It used something called whole-genome shotgun sequencing.
Revolutionizing sequencing using the whole genome
Instead of relying on restriction enzymes, whole genome shotgun sequencing randomly chops the genome into fragments and relies on computer modeling to reconstruct the whole sequence. This requires an enormous amount of processing power and very sophisticated computer models to sift through the data. The amount of computing power needed would not have been possible in the late ‘80s when the project started. The late start meant that both the Human Genome Project and Celera completed their sequence at roughly the same time.
Today, the digital revolution has brought whole genome shotgun sequencing to the forefront. The ease with which one can dump an entire genome into an algorithm and get a completed, accurate sequence far surpasses the tedious Sanger Sequencing of the original project.
Completing the genome sequence
The final draft of the sequence was announced on April 14, 2003. The sequence covered roughly 92% of the genome and both systems corroborated the results. From the start, the goal of the project excluded hard-to-read areas, such as end telomeres and connection points, as well as the Y chromosome. These have later been sequenced, and gaps in the genome have been identified over the years. As they have been identified and corrected, newer versions of the sequence have been released, but the majority of it has remained unchanged. This has revolutionized our ability to identify mutations, identify causes of cancers, and, of course, help genealogists make connections.
Honoring scientific achievement and progress on DNA Day
If you have made it this far, congratulations. I award you fourteen gold stars, one for each section. It’s been a journey through incellption and back. Yes, I am milking that term for all it is worth. I hope that now when you see a double helix, you won’t just shrug it off as just a pretty shape, but recognize the countless hours of research and intuitive leaps that brought it from cork under a magnifying glass to the cultural icon and revolutionary contribution to science that we know today.
DNA Day was first celebrated in 2003 to coincide with the final release of the human genome, and April 25 was chosen as a fitting day to honor the momentous paper published on that day in 1953. To me, it celebrates the human pyramid of geniuses standing on the shoulders of geniuses to bring us to where we are now. Where do you go from here? You can explore the wonderful world of your own chromosomes with a DNA test!

About the Author
Jim Brewster
Subject Matter Expert at FamilyTreeDNA
Jim Brewster was born at a very early age and gradually became older. He has been in the genetic genealogy field since 2014 and delivered numerous presentations at genealogy conferences. He has helped with collaborations between FamilyTreeDNA and non-profit organizations and for some reason FamilyTreeDNA decided to let him write stuff too.
With a proven track record of both doing things and accomplishing stuff, Jim enjoys presenting and writing about genetic genealogy methods and the science of DNA testing. In his free time, he enjoys puns and cat pictures.